
![]()
Molecular Modeling and Structure-based Drug Design Technologies
PIEFII patent rights
- The Ultimate Technology In The Drug Discovery -
Institute of Molecular Function now announces the development of the drug discovery technology in the next-generation. This novel technology will serve as the core technology for the structure-based drug design.
The PIEFII is an acronym formed from the Potential Interaction Energy Field with Interaction Information.
PIEFII technology can precisely predict the ligand-binding site for any kinds of proteins using only the three-dimensional coordinate of the whole structure as well as can visualize the scaffold and the pharmacophores of a drug candidate at the positions of the potential heavy atoms in the predicted ligand-binding site of the target protein. Thus, this technology can also support the logical molecular modeling for a target protein in which the ligand-binding site has never been identified. The researchers can select the drug candidates of the target protein from only their knowledge and experiences. In the drug design, the predicted information on the potential heavy atoms serves as the directionality of the molecular modeling such as the lead optimizations.
The PIEFII technology, which is the core technology for performing the molecular superposition, is carried on the Docking Study with HyperChem and the Virtual Screening System and thus supporting the efficient searches of a drug candidate from an arbitrary chemical database.
The following figures show the results of the prediction of positions at which the potential heavy-atoms in an interaction partner will stabilize with the minimum interaction energy. The right-hand side of the figures shows the results given from mapping the predicted chemical properties on the above predicted heavy-atoms and simultaneously shows the complexed natural ligand. The predicted ligand-binding site containing these potential heavy-atoms was perfectly identical with the true ligand-binding site. Moreover, the three-dimensional scaffold or skeleton constructed from the individual potential heavy-atoms was identical to that of the natural ligand. The predicted chemical properties on the potential heavy-atoms excellently reproduced those of the natural ligand, as well. In particular, the results well reproduced the pharmacophore corresponding to the carboxylic acid of the natural ligand.
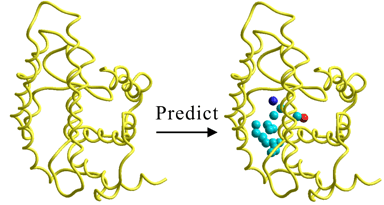
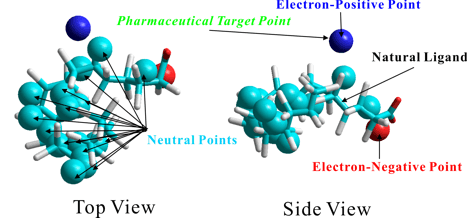
The following figure of the left-hand side shows the results of the prediction for the apo-form protein which is not complexed with any ligands. The figure of the right-hand side shows the results of the prediction for the corresponding holo-form protein which is complexed with its natural ligand. As the results of comparison between both figures, it is apparent that the PIEFII technology can precisely predict the potential ligand-binding site of the apo-form protein which is stabilized by itself. On the other hand, the both results suggested importance of the internal structure for the holo-form formation processes. That is, the Phe residue of the holo-form moves to the upper site of the predicted binding site of the apo-form, thus giving the true binding site to the holo-form. The predicted ligand-binding site of the holo-form protein was perfectly identical with the true ligand-binding site. Moreover, the three-dimensional scaffold or skeleton constructed from the individual potential heavy-atoms was identical to that of the natural ligand. The predicted chemical properties on the potential heavy-atoms excellently reproduced those of the natural ligand, as well.

See other results.
Refer to an objective opinion.
Our customers are very interested in the PIEFII technology.
A new logical drug design method using the PIEFII technology (Japanese)













